Import Data
For this example I am using the PoliticalDemocracy data set from the lavaan package.
Descriptives
Descriptive Statistics
|
Variable
|
n
|
Mean
|
SD
|
min
|
max
|
Skewness
|
Kurtosis
|
% Missing
|
|
x1
|
75
|
5.05
|
0.73
|
3.78
|
6.74
|
0.26
|
-0.69
|
0
|
|
x2
|
75
|
4.79
|
1.51
|
1.39
|
7.87
|
-0.36
|
-0.50
|
0
|
|
x3
|
75
|
3.56
|
1.41
|
1.00
|
6.42
|
0.09
|
-0.88
|
0
|
|
y1
|
75
|
5.46
|
2.62
|
1.25
|
10.00
|
-0.10
|
-1.10
|
0
|
|
y2
|
75
|
4.26
|
3.95
|
0.00
|
10.00
|
0.33
|
-1.43
|
0
|
|
y3
|
75
|
6.56
|
3.28
|
0.00
|
10.00
|
-0.62
|
-0.66
|
0
|
|
y4
|
75
|
4.45
|
3.35
|
0.00
|
10.00
|
0.12
|
-1.16
|
0
|
|
y5
|
75
|
5.14
|
2.61
|
0.00
|
10.00
|
-0.24
|
-0.72
|
0
|
|
y6
|
75
|
2.98
|
3.37
|
0.00
|
10.00
|
0.93
|
-0.40
|
0
|
|
y7
|
75
|
6.20
|
3.29
|
0.00
|
10.00
|
-0.58
|
-0.67
|
0
|
|
y8
|
75
|
4.04
|
3.25
|
0.00
|
10.00
|
0.46
|
-0.91
|
0
|
Correlation Matrix
|
|
y1
|
y2
|
y3
|
y4
|
y5
|
y6
|
y7
|
y8
|
x1
|
x2
|
x3
|
|
y1
|
|
0.60***
|
0.68***
|
0.69***
|
0.74***
|
0.65***
|
0.67***
|
0.67***
|
0.38***
|
0.32**
|
0.25*
|
|
y2
|
0.60***
|
|
0.45***
|
0.72***
|
0.54***
|
0.71***
|
0.58***
|
0.61***
|
0.21
|
0.25*
|
0.21
|
|
y3
|
0.68***
|
0.45***
|
|
0.61***
|
0.58***
|
0.43***
|
0.65***
|
0.53***
|
0.33**
|
0.31**
|
0.23
|
|
y4
|
0.69***
|
0.72***
|
0.61***
|
|
0.65***
|
0.66***
|
0.68***
|
0.74***
|
0.47***
|
0.44***
|
0.39***
|
|
y5
|
0.74***
|
0.54***
|
0.58***
|
0.65***
|
|
0.56***
|
0.68***
|
0.63***
|
0.56***
|
0.52***
|
0.43***
|
|
y6
|
0.65***
|
0.71***
|
0.43***
|
0.66***
|
0.56***
|
|
0.61***
|
0.75***
|
0.34**
|
0.35**
|
0.33**
|
|
y7
|
0.67***
|
0.58***
|
0.65***
|
0.68***
|
0.68***
|
0.61***
|
|
0.71***
|
0.39***
|
0.40***
|
0.35**
|
|
y8
|
0.67***
|
0.61***
|
0.53***
|
0.74***
|
0.63***
|
0.75***
|
0.71***
|
|
0.46***
|
0.46***
|
0.37**
|
|
x1
|
0.38***
|
0.21
|
0.33**
|
0.47***
|
0.56***
|
0.34**
|
0.39***
|
0.46***
|
|
0.89***
|
0.80***
|
|
x2
|
0.32**
|
0.25*
|
0.31**
|
0.44***
|
0.52***
|
0.35**
|
0.40***
|
0.46***
|
0.89***
|
|
0.85***
|
|
x3
|
0.25*
|
0.21
|
0.23
|
0.39***
|
0.43***
|
0.33**
|
0.35**
|
0.37**
|
0.80***
|
0.85***
|
|
|
Computed correlation used pearson-method with pairwise-deletion.
|
CFA
Summary Output
Model Significance
|
Sample.Size
|
Chi.Square
|
df
|
p.value
|
|
75
|
38.125
|
35
|
0.329
|
Model Fit Measures
|
CFI
|
RMSEA
|
RMSEA.Lower
|
RMSEA.Upper
|
AIC
|
BIC
|
|
0.995
|
0.035
|
0
|
0.092
|
3179.582
|
3276.916
|
Factor Loadings
|
|
|
Loadings
|
|
|
Latent Factor
|
Indicator
|
Standardized
|
Unstandardized
|
SE
|
z
|
sig
|
|
ind60
|
x1
|
0.920
|
0.670
|
0.065
|
10.339
|
***
|
|
ind60
|
x2
|
0.973
|
1.460
|
0.128
|
11.423
|
***
|
|
ind60
|
x3
|
0.872
|
1.218
|
0.128
|
9.489
|
***
|
|
dem60
|
y1
|
0.850
|
2.223
|
0.253
|
8.789
|
***
|
|
dem60
|
y2
|
0.717
|
2.794
|
0.408
|
6.850
|
***
|
|
dem60
|
y3
|
0.722
|
2.351
|
0.341
|
6.902
|
***
|
|
dem60
|
y4
|
0.846
|
2.812
|
0.324
|
8.677
|
***
|
|
dem65
|
y5
|
0.808
|
2.103
|
0.256
|
8.210
|
***
|
|
dem65
|
y6
|
0.746
|
2.493
|
0.341
|
7.320
|
***
|
|
dem65
|
y7
|
0.824
|
2.691
|
0.319
|
8.441
|
***
|
|
dem65
|
y8
|
0.828
|
2.662
|
0.316
|
8.430
|
***
|
Latent Factor Correlations
|
Factor 1
|
Factor 2
|
r
|
sig
|
|
ind60
|
dem60
|
0.447
|
***
|
|
ind60
|
dem65
|
0.578
|
***
|
|
dem60
|
dem65
|
0.967
|
***
|
Latent Factor Variance/Residual Variance
|
Factor 1
|
Factor 2
|
var
|
var.std
|
sig
|
|
R-Squared Values
|
Variable
|
R-Squared
|
|
x1
|
0.8461294
|
|
x2
|
0.9467924
|
|
x3
|
0.7606256
|
|
y1
|
0.7232242
|
|
y2
|
0.5142640
|
|
y3
|
0.5217883
|
|
y4
|
0.7152245
|
|
y5
|
0.6528920
|
|
y6
|
0.5565270
|
|
y7
|
0.6784378
|
|
y8
|
0.6853215
|
Diagram Output
semPaths(fit, whatLabels = "std", edge.label.cex = .5, layout = "tree2", rotation = 2, style = "lisrel", intercepts = FALSE, residuals = TRUE, curve = 1, curvature = 3, nCharNodes = 8, sizeMan = 6, sizeMan2 = 3, optimizeLatRes = TRUE, edge.color = "#000000", latents = factors)
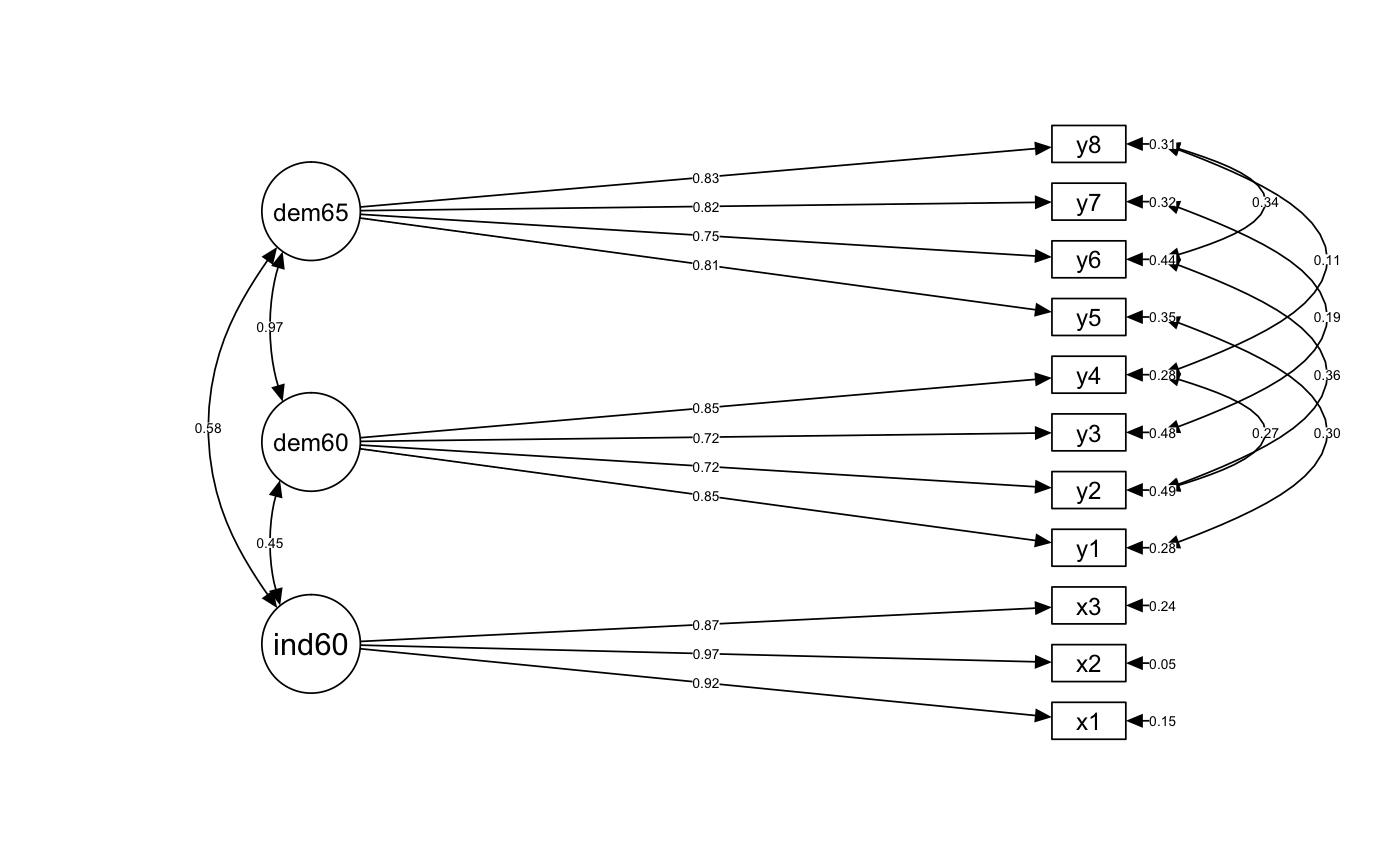
Residual Correlation Matrix
|
|
x1
|
x2
|
x3
|
y1
|
y2
|
y3
|
y4
|
y5
|
y6
|
y7
|
y8
|
|
x1
|
|
|
|
|
|
|
|
|
|
|
|
|
x2
|
0.00
|
|
|
|
|
|
|
|
|
|
|
|
x3
|
0.00
|
0.00
|
|
|
|
|
|
|
|
|
|
|
y1
|
0.03
|
-0.05
|
-0.08
|
|
|
|
|
|
|
|
|
|
y2
|
-0.08
|
-0.06
|
-0.07
|
-0.01
|
|
|
|
|
|
|
|
|
y3
|
0.03
|
0.00
|
-0.06
|
0.06
|
-0.07
|
|
|
|
|
|
|
|
y4
|
0.12
|
0.08
|
0.06
|
-0.03
|
0.01
|
0.00
|
|
|
|
|
|
|
y5
|
0.14
|
0.07
|
0.02
|
-0.02
|
-0.02
|
0.01
|
-0.01
|
|
|
|
|
|
y6
|
-0.05
|
-0.06
|
-0.04
|
0.04
|
0.02
|
-0.09
|
0.05
|
-0.04
|
|
|
|
|
y7
|
-0.05
|
-0.06
|
-0.06
|
0.00
|
0.01
|
0.00
|
0.01
|
0.01
|
-0.01
|
|
|
|
y8
|
0.02
|
-0.01
|
-0.05
|
-0.01
|
0.03
|
-0.05
|
0.03
|
-0.04
|
0.01
|
0.03
|
|
Full Output
## lavaan 0.6-3 ended normally after 75 iterations
##
## Optimization method NLMINB
## Number of free parameters 42
##
## Number of observations 75
## Number of missing patterns 1
##
## Estimator ML
## Model Fit Test Statistic 38.125
## Degrees of freedom 35
## P-value (Chi-square) 0.329
##
## Model test baseline model:
##
## Minimum Function Test Statistic 730.654
## Degrees of freedom 55
## P-value 0.000
##
## User model versus baseline model:
##
## Comparative Fit Index (CFI) 0.995
## Tucker-Lewis Index (TLI) 0.993
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1547.791
## Loglikelihood unrestricted model (H1) -1528.728
##
## Number of free parameters 42
## Akaike (AIC) 3179.582
## Bayesian (BIC) 3276.916
## Sample-size adjusted Bayesian (BIC) 3144.543
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.035
## 90 Percent Confidence Interval 0.000 0.092
## P-value RMSEA <= 0.05 0.611
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.041
##
## Parameter Estimates:
##
## Information Observed
## Observed information based on Hessian
## Standard Errors Standard
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## ind60 =~
## x1 0.670 0.065 10.339 0.000 0.670 0.920
## x2 1.460 0.128 11.423 0.000 1.460 0.973
## x3 1.218 0.128 9.489 0.000 1.218 0.872
## dem60 =~
## y1 2.223 0.253 8.789 0.000 2.223 0.850
## y2 2.794 0.408 6.850 0.000 2.794 0.717
## y3 2.351 0.341 6.902 0.000 2.351 0.722
## y4 2.812 0.324 8.677 0.000 2.812 0.846
## dem65 =~
## y5 2.103 0.256 8.210 0.000 2.103 0.808
## y6 2.493 0.341 7.320 0.000 2.493 0.746
## y7 2.691 0.319 8.441 0.000 2.691 0.824
## y8 2.662 0.316 8.430 0.000 2.662 0.828
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .y1 ~~
## .y5 0.624 0.369 1.690 0.091 0.624 0.296
## .y2 ~~
## .y4 1.313 0.699 1.879 0.060 1.313 0.273
## .y6 2.153 0.726 2.964 0.003 2.153 0.356
## .y3 ~~
## .y7 0.795 0.621 1.280 0.201 0.795 0.191
## .y4 ~~
## .y8 0.348 0.458 0.761 0.447 0.348 0.109
## .y6 ~~
## .y8 1.356 0.572 2.371 0.018 1.356 0.338
## ind60 ~~
## dem60 0.447 0.105 4.267 0.000 0.447 0.447
## dem65 0.578 0.089 6.466 0.000 0.578 0.578
## dem60 ~~
## dem65 0.967 0.029 32.840 0.000 0.967 0.967
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .x1 5.054 0.084 60.127 0.000 5.054 6.943
## .x2 4.792 0.173 27.657 0.000 4.792 3.194
## .x3 3.558 0.161 22.066 0.000 3.558 2.548
## .y1 5.465 0.302 18.104 0.000 5.465 2.090
## .y2 4.256 0.450 9.461 0.000 4.256 1.093
## .y3 6.563 0.376 17.460 0.000 6.563 2.016
## .y4 4.453 0.384 11.598 0.000 4.453 1.339
## .y5 5.136 0.301 17.092 0.000 5.136 1.974
## .y6 2.978 0.386 7.717 0.000 2.978 0.891
## .y7 6.196 0.377 16.427 0.000 6.196 1.897
## .y8 4.043 0.371 10.889 0.000 4.043 1.257
## ind60 0.000 0.000 0.000
## dem60 0.000 0.000 0.000
## dem65 0.000 0.000 0.000
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .x1 0.082 0.020 4.136 0.000 0.082 0.154
## .x2 0.120 0.070 1.712 0.087 0.120 0.053
## .x3 0.467 0.089 5.233 0.000 0.467 0.239
## .y1 1.891 0.469 4.035 0.000 1.891 0.277
## .y2 7.373 1.346 5.479 0.000 7.373 0.486
## .y3 5.067 0.968 5.233 0.000 5.067 0.478
## .y4 3.148 0.756 4.165 0.000 3.148 0.285
## .y5 2.351 0.489 4.810 0.000 2.351 0.347
## .y6 4.954 0.895 5.532 0.000 4.954 0.443
## .y7 3.431 0.728 4.715 0.000 3.431 0.322
## .y8 3.254 0.707 4.603 0.000 3.254 0.315
## ind60 1.000 1.000 1.000
## dem60 1.000 1.000 1.000
## dem65 1.000 1.000 1.000
## name idx nobs type exo user mean var nlev lnam
## 1 x1 9 75 numeric 0 0 5.054 0.537 0
## 2 x2 10 75 numeric 0 0 4.792 2.282 0
## 3 x3 11 75 numeric 0 0 3.558 1.976 0
## 4 y1 1 75 numeric 0 0 5.465 6.879 0
## 5 y2 2 75 numeric 0 0 4.256 15.580 0
## 6 y3 3 75 numeric 0 0 6.563 10.764 0
## 7 y4 4 75 numeric 0 0 4.453 11.219 0
## 8 y5 5 75 numeric 0 0 5.136 6.826 0
## 9 y6 6 75 numeric 0 0 2.978 11.375 0
## 10 y7 7 75 numeric 0 0 6.196 10.799 0
## 11 y8 8 75 numeric 0 0 4.043 10.534 0
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 52 ind60 =~ y4 4.796 0.577 0.577 0.174 0.174
## 53 ind60 =~ y5 4.456 0.559 0.559 0.215 0.215
## 70 dem65 =~ y4 4.260 2.986 2.986 0.898 0.898
## 99 y1 ~~ y3 3.771 0.849 0.849 0.274 0.274
## 74 x1 ~~ y2 3.040 -0.155 -0.155 -0.200 -0.200
SEM
Summary Output
Model Significance
|
Sample.Size
|
Chi.Square
|
df
|
p.value
|
|
75
|
37.617
|
35
|
0.35
|
Model Fit Measures
|
CFI
|
RMSEA
|
RMSEA.Lower
|
RMSEA.Upper
|
AIC
|
BIC
|
|
0.996
|
0.032
|
0
|
0.091
|
3179.582
|
3276.916
|
Factor Loadings
|
|
|
Loadings
|
|
|
Latent Factor
|
Indicator
|
Standardized
|
Unstandardized
|
SE
|
z
|
sig
|
|
ind60
|
x1
|
0.920
|
1.000
|
0.000
|
|
|
|
ind60
|
x2
|
0.973
|
2.180
|
0.140
|
15.580
|
***
|
|
ind60
|
x3
|
0.872
|
1.819
|
0.153
|
11.869
|
***
|
|
dem60
|
y1
|
0.850
|
1.000
|
0.000
|
|
|
|
dem60
|
y2
|
0.717
|
1.257
|
0.187
|
6.730
|
***
|
|
dem60
|
y3
|
0.722
|
1.058
|
0.149
|
7.083
|
***
|
|
dem60
|
y4
|
0.846
|
1.265
|
0.152
|
8.335
|
***
|
|
dem65
|
y5
|
0.808
|
1.000
|
0.000
|
|
|
|
dem65
|
y6
|
0.746
|
1.186
|
0.173
|
6.873
|
***
|
|
dem65
|
y7
|
0.824
|
1.280
|
0.161
|
7.925
|
***
|
|
dem65
|
y8
|
0.828
|
1.266
|
0.164
|
7.704
|
***
|
Regression Paths
|
DV
|
Predictor
|
Beta
|
B
|
SE
|
z
|
sig
|
|
dem60
|
ind60
|
0.447
|
1.483
|
0.400
|
3.708
|
***
|
|
dem65
|
ind60
|
0.182
|
0.572
|
0.235
|
2.432
|
|
|
dem65
|
dem60
|
0.885
|
0.837
|
0.099
|
8.419
|
***
|
Latent Factor Correlations
|
Factor 1
|
Factor 2
|
r
|
sig
|
|
R-Squared Values
|
Variable
|
R-Squared
|
|
x1
|
0.8461294
|
|
x2
|
0.9467924
|
|
x3
|
0.7606258
|
|
y1
|
0.7232244
|
|
y2
|
0.5142639
|
|
y3
|
0.5217888
|
|
y4
|
0.7152247
|
|
y5
|
0.6528919
|
|
y6
|
0.5565269
|
|
y7
|
0.6784379
|
|
y8
|
0.6853214
|
|
dem60
|
0.1995524
|
|
dem65
|
0.9609950
|
Diagram Output
semPaths(fit, whatLabels = "std", edge.label.cex = .5, layout = "tree2", rotation = 2, style = "lisrel", intercepts = FALSE, residuals = TRUE, curve = 1, curvature = 3, nCharNodes = 8, sizeMan = 6, sizeMan2 = 3, optimizeLatRes = TRUE, edge.color = "#000000", latents = factors)
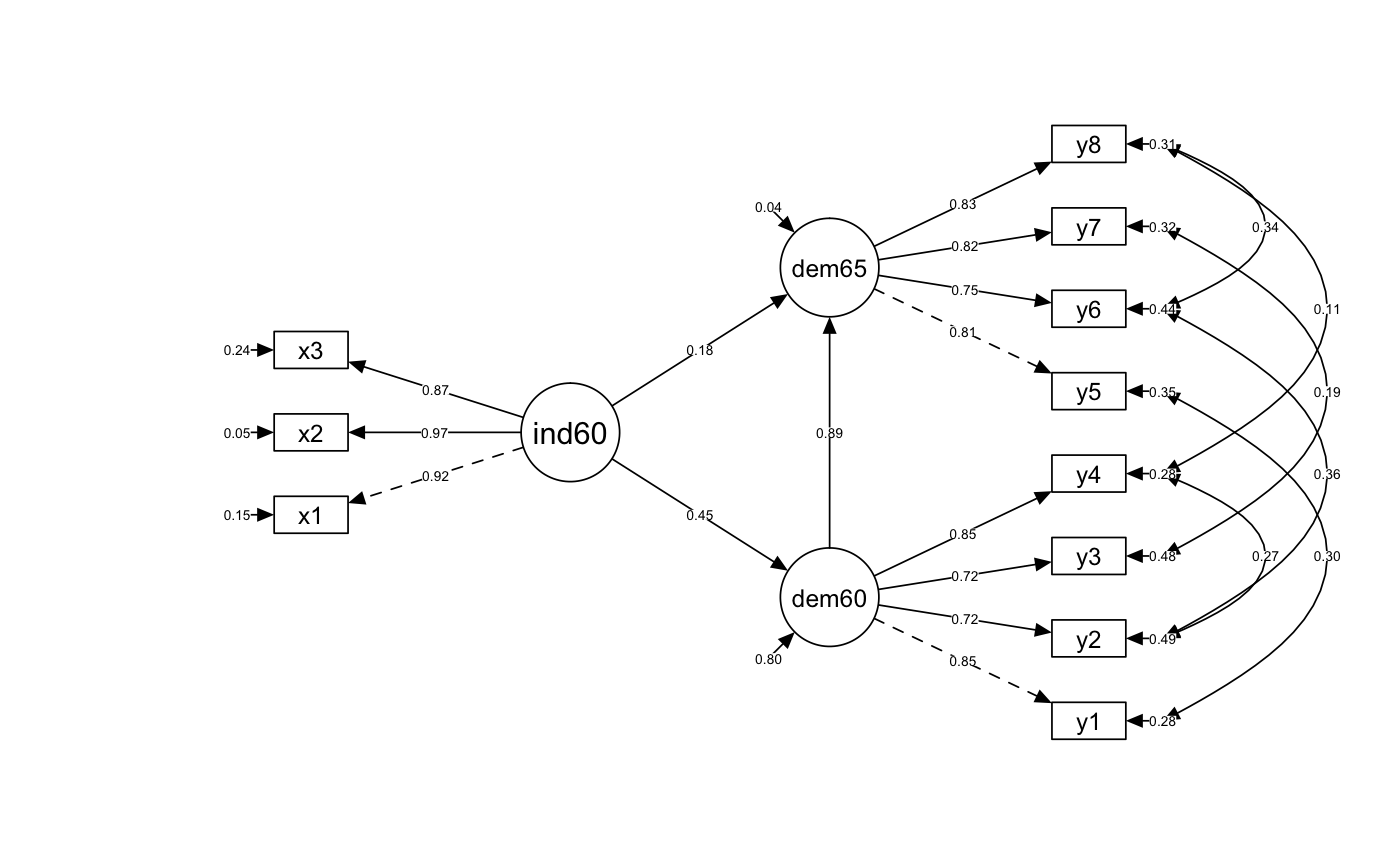
Residual Correlation Matrix
|
|
x1
|
x2
|
x3
|
y1
|
y2
|
y3
|
y4
|
y5
|
y6
|
y7
|
y8
|
|
x1
|
|
|
|
|
|
|
|
|
|
|
|
|
x2
|
0.00
|
|
|
|
|
|
|
|
|
|
|
|
x3
|
0.00
|
0.00
|
|
|
|
|
|
|
|
|
|
|
y1
|
0.03
|
-0.05
|
-0.09
|
|
|
|
|
|
|
|
|
|
y2
|
-0.08
|
-0.06
|
-0.07
|
0.00
|
|
|
|
|
|
|
|
|
y3
|
0.03
|
0.00
|
-0.06
|
0.06
|
-0.06
|
|
|
|
|
|
|
|
y4
|
0.12
|
0.08
|
0.06
|
-0.03
|
0.02
|
0.00
|
|
|
|
|
|
|
y5
|
0.13
|
0.07
|
0.02
|
-0.02
|
-0.01
|
0.01
|
-0.01
|
|
|
|
|
|
y6
|
-0.05
|
-0.06
|
-0.04
|
0.04
|
0.03
|
-0.09
|
0.05
|
-0.04
|
|
|
|
|
y7
|
-0.05
|
-0.06
|
-0.06
|
-0.01
|
0.01
|
0.00
|
0.01
|
0.01
|
0.00
|
|
|
|
y8
|
0.02
|
-0.01
|
-0.05
|
-0.01
|
0.04
|
-0.05
|
0.03
|
-0.04
|
0.01
|
0.03
|
|
Full Output
## lavaan 0.6-3 ended normally after 83 iterations
##
## Optimization method NLMINB
## Number of free parameters 42
##
## Number of observations 75
## Number of missing patterns 1
##
## Estimator ML
## Model Fit Test Statistic 37.617
## Degrees of freedom 35
## P-value (Chi-square) 0.350
##
## Model test baseline model:
##
## Minimum Function Test Statistic 720.912
## Degrees of freedom 55
## P-value 0.000
##
## User model versus baseline model:
##
## Comparative Fit Index (CFI) 0.996
## Tucker-Lewis Index (TLI) 0.994
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1547.791
## Loglikelihood unrestricted model (H1) -1528.728
##
## Number of free parameters 42
## Akaike (AIC) 3179.582
## Bayesian (BIC) 3276.916
## Sample-size adjusted Bayesian (BIC) 3144.543
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.032
## 90 Percent Confidence Interval 0.000 0.091
## P-value RMSEA <= 0.05 0.629
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.041
##
## Parameter Estimates:
##
## Information Observed
## Observed information based on Hessian
## Standard Errors Standard
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## ind60 =~
## x1 1.000 0.670 0.920
## x2 2.180 0.140 15.580 0.000 1.460 0.973
## x3 1.819 0.153 11.869 0.000 1.218 0.872
## dem60 =~
## y1 1.000 2.223 0.850
## y2 1.257 0.187 6.730 0.000 2.794 0.717
## y3 1.058 0.149 7.083 0.000 2.351 0.722
## y4 1.265 0.152 8.335 0.000 2.812 0.846
## dem65 =~
## y5 1.000 2.103 0.808
## y6 1.186 0.173 6.873 0.000 2.493 0.746
## y7 1.280 0.161 7.925 0.000 2.691 0.824
## y8 1.266 0.164 7.704 0.000 2.662 0.828
##
## Regressions:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## dem60 ~
## ind60 1.483 0.400 3.708 0.000 0.447 0.447
## dem65 ~
## ind60 0.572 0.235 2.432 0.015 0.182 0.182
## dem60 0.837 0.099 8.419 0.000 0.885 0.885
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .y1 ~~
## .y5 0.624 0.371 1.679 0.093 0.624 0.296
## .y2 ~~
## .y4 1.313 0.703 1.867 0.062 1.313 0.273
## .y6 2.153 0.731 2.944 0.003 2.153 0.356
## .y3 ~~
## .y7 0.795 0.625 1.271 0.204 0.795 0.191
## .y4 ~~
## .y8 0.348 0.461 0.755 0.450 0.348 0.109
## .y6 ~~
## .y8 1.356 0.576 2.355 0.019 1.356 0.338
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .x1 5.054 0.085 59.724 0.000 5.054 6.943
## .x2 4.792 0.174 27.472 0.000 4.792 3.194
## .x3 3.558 0.162 21.918 0.000 3.558 2.548
## .y1 5.465 0.304 17.983 0.000 5.465 2.090
## .y2 4.256 0.453 9.398 0.000 4.256 1.093
## .y3 6.563 0.378 17.344 0.000 6.563 2.016
## .y4 4.453 0.386 11.520 0.000 4.453 1.339
## .y5 5.136 0.303 16.977 0.000 5.136 1.974
## .y6 2.978 0.389 7.665 0.000 2.978 0.891
## .y7 6.196 0.380 16.317 0.000 6.196 1.897
## .y8 4.043 0.374 10.816 0.000 4.043 1.257
## ind60 0.000 0.000 0.000
## .dem60 0.000 0.000 0.000
## .dem65 0.000 0.000 0.000
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .x1 0.082 0.020 4.109 0.000 0.082 0.154
## .x2 0.120 0.070 1.701 0.089 0.120 0.053
## .x3 0.467 0.090 5.198 0.000 0.467 0.239
## .y1 1.891 0.472 4.008 0.000 1.891 0.277
## .y2 7.373 1.355 5.442 0.000 7.373 0.486
## .y3 5.067 0.975 5.198 0.000 5.067 0.478
## .y4 3.148 0.761 4.137 0.000 3.148 0.285
## .y5 2.351 0.492 4.778 0.000 2.351 0.347
## .y6 4.954 0.901 5.495 0.000 4.954 0.443
## .y7 3.431 0.733 4.683 0.000 3.431 0.322
## .y8 3.254 0.712 4.572 0.000 3.254 0.315
## ind60 0.448 0.087 5.135 0.000 1.000 1.000
## .dem60 3.956 0.951 4.160 0.000 0.800 0.800
## .dem65 0.172 0.222 0.778 0.437 0.039 0.039
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 52 ind60 =~ y4 4.796 0.862 0.577 0.174 0.174
## 53 ind60 =~ y5 4.456 0.835 0.559 0.215 0.215
## 70 dem65 =~ y4 4.260 1.420 2.986 0.898 0.898
## 99 y1 ~~ y3 3.771 0.849 0.849 0.274 0.274
## 74 x1 ~~ y2 3.040 -0.155 -0.155 -0.200 -0.200

